Speeding up Python with Mojo🔥
Speedup Python implementation in Mojo for simple Euclidean distance calculation
In this example we’ll calculate the Euclidean distance between two n-dimensional vectors a and b mathematically expressed as the L2-norm of the difference vector: $$ ||a-b||_2 $$
Let’s start by creating 2 random n-dimensional numpy arrays in python
Notice that the cell starts with %%python i.e. this cell will execute in Python, no Mojo. We’ll use this to compare performance between Python and Mojo implementations.
%%python
import time
import numpy as np
from math import sqrt
from timeit import timeit
n = 1000000
np.random.seed(42)
arr1_np = np.random.rand(n)
arr2_np = np.random.rand(n)
arr1_list = arr1_np.tolist()
arr2_list = arr2_np.tolist()
def print_formatter(string, value):
print(f"{string}: {value}")
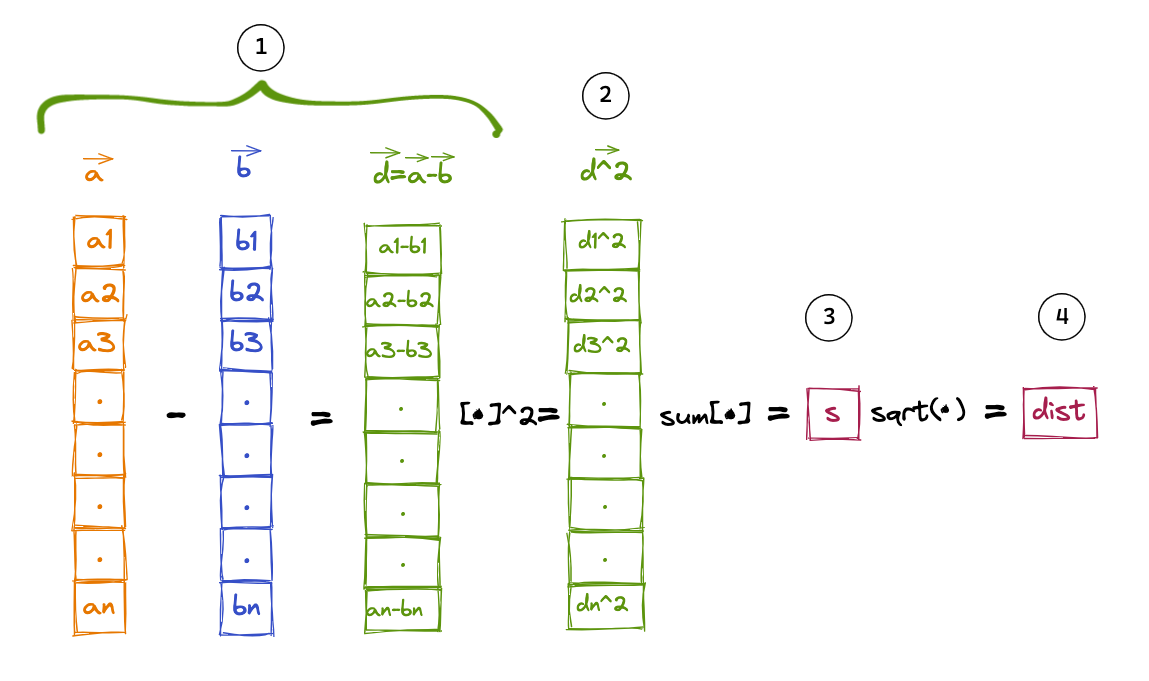
Below is a Python function which takes arr1_list and arr2_list as arguments and calculates the eculidean distance using the following steps:
- Calculate the element-wise difference between two vectors to create a difference vector
- Square each element in the difference vector
- Sum up all the squared elements of the difference vector
- Take the square root of the sum
Go ahead and run the cell and take a note of the value and duration
%%python
# Pure python iterative implementation
def python_dist(a,b):
sq_dist = 0.0
n = len(a)
for i in range(n):
diff = a[i]-b[i]
sq_dist += diff*diff
return sqrt(sq_dist)
tm = timeit(lambda: python_dist(arr1_list,arr2_list), number=5)/5
print("=== Pure Python Performance ===")
print_formatter("value", python_dist(arr1_list,arr2_list))
print_formatter("time (ms)", 1000*tm)
Let’s compare pure-Python performance with NumPy. NumPy takes advantage of optimized linear algebra subroutines to accelerate these calculations. Run the cell below and take a note of the speedup.
%%python
def python_numpy_dist(a,b):
return np.linalg.norm(a-b)
tm = timeit(lambda: python_numpy_dist(arr1_np,arr2_np), number=5)/5
print("=== NumPy Performance ===")
print_formatter("value", python_numpy_dist(arr1_np,arr2_np))
print_formatter("time (ms)", 1000*tm)
Time to Mojo🔥! Run the following cell to copy over the numpy arrays from Python to Mojo🔥 tensors so can compare not only the execution time but also verify the accuracy of the calculations.
from tensor import Tensor
from time import now
from math import sqrt
let n: Int = 1000000
alias dtype = DType.float64
var arr1_tensor = Tensor[dtype](n)
var arr2_tensor = Tensor[dtype](n)
for i in range(n):
arr1_tensor[i] = arr1_np[i].to_float64()
arr2_tensor[i] = arr2_np[i].to_float64()
Exercise: Copy over the pure-Python function into Mojo🔥 and variable declaration and return types, you did in notebook 1.
Click for solution ✅
fn mojo_dist(a: Tensor[dtype], b: Tensor[dtype]) -> Float64:
var sq_dist:Float64 = 0.0
let n = a.num_elements()
for i in range(n):
let diff = a[i]-b[i]
sq_dist += diff*diff
return sqrt(sq_dist)
#Hint: Insert function below with the following signature
#fn mojo_dist(a: Tensor[dtype], b: Tensor[dtype]) -> Float64: ...
Copy paste the following the same cell to benchmark
let eval_begin = now()
let mojo_arr_sum = mojo_dist(arr1_tensor,arr2_tensor)
let eval_end = now()
print("=== Mojo Performance ===")
print_formatter("value", mojo_arr_sum)
print_formatter("time (ms)", Float64((eval_end - eval_begin)) / 1e6)
Accelerating Mojo🔥 code with Vectorization
Modern CPU cores have dedicated vector register that can perform calculations simultaneously on fixed length vector data. For example an Intel CPU with AVX 512 has 512-bit vector register, therefore we have 512/64=8 Float64 elements on which we can simultaneously perform calculations. This is referred to as SIMD = Single Instruction Multiple Data. The theoretical speed up is 8x but in practice it’ll be lower due to memory reads/writes latency.
Note: SIMD is should not be confused with parallel processing with multiple cores/threads. Each core has dedicated SIMD vector units and we can take advantage of both SIMD and parallel processing to see massive speedup as you’ll see in notebook 3
First, let’s gather the simd_width for specific dtype on the specific CPU you’ll be running this on.
from sys.info import simdwidthof
from algorithm import vectorize
alias simd_width = simdwidthof[DType.float64]()
To vectorize our naive mojo_dist function, we’ll write a closure that is parameterized on simd_width. Rather than operate on individual elements, we’ll work with simd_width elements which gives us speed ups. You can access simd_width elements using simd_load.
Given that this is a complex topic, I’ve provided the solution below, please raise your hand and ask the instructor for explainations if anything is unclear.
fn mojo_dist_vectorized(a: Tensor[DType.float64], b: Tensor[DType.float64]) -> Float64:
var sq_dist: Float64 = 0.0
@parameter
fn simd_norm[simd_width:Int](idx:Int):
let diff = a.simd_load[simd_width](idx) - b.simd_load[simd_width](idx)
sq_dist += (diff*diff).reduce_add()
vectorize[simd_width, simd_norm](a.num_elements())
return sqrt(sq_dist)
let eval_begin = now()
let mojo_arr_vec_sum = mojo_dist_vectorized(arr1_tensor,arr2_tensor)
let eval_end = now()
print_formatter("value", mojo_arr_vec_sum)
print_formatter("time (ms)",((eval_end - eval_begin)) / 1e6)